Research Overview
Our laboratory combines traditional neuropathology methods with quantitative approaches to advance brain science in the areas of brain aging, cancer, neurotrauma, and neurodegenerative disease.
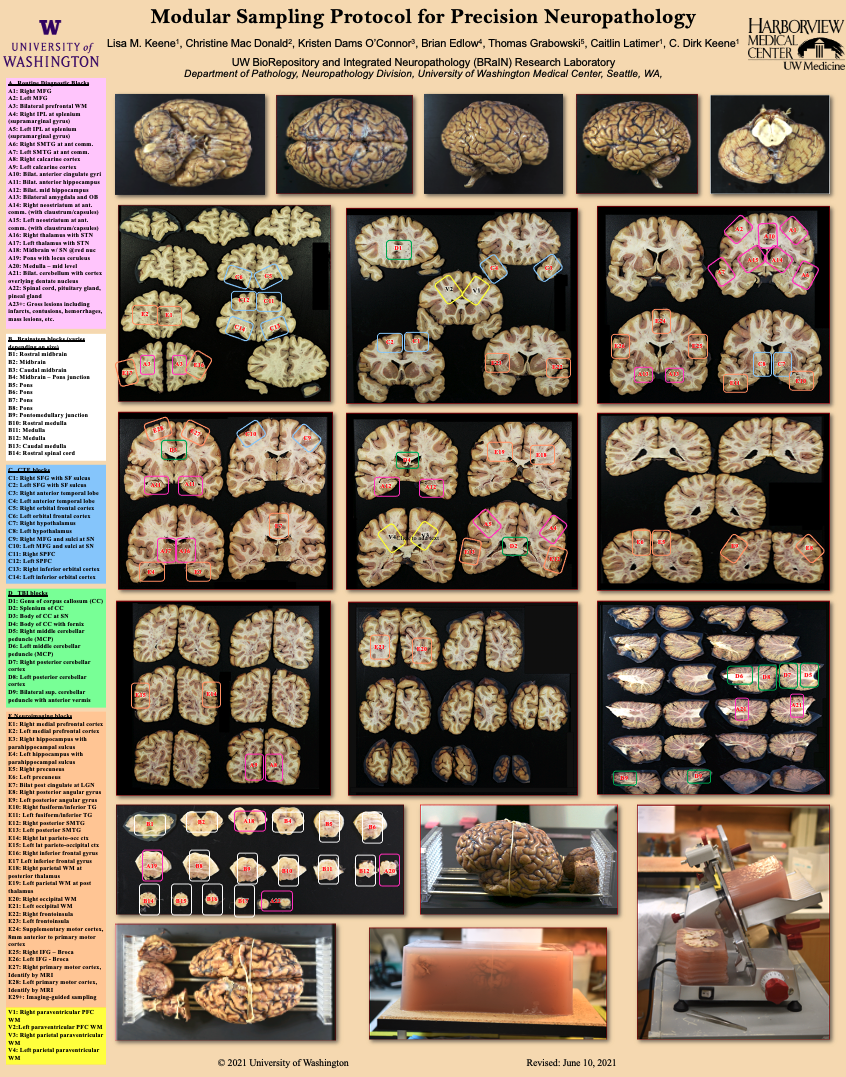
Rapid Brain Removal & Brain Research Report
Timely acquisition and preservation of brain and spinal cord tissue is essential to the BRaIN lab’s research and tissue sharing efforts. Our dedicated brain removal and research team facilitates the generous gifts of over 200 donors each year in addition to maintaining a biorepository of fresh frozen and formalin-fixed specimens from thousands of subjects.
Ex-Vivo MRI & 3D Image Reconstructions
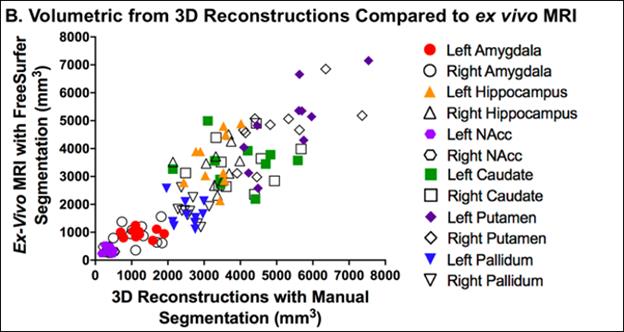
Ex-vivo MRI of fixed brain tissue, led by UW Professor of Neurological Surgery Dr. Christine Mac Donald, provides an important tool to guide regional sampling for neuropathologic evaluation and measure volumes of diverse structures across the brain. Combined with MRI imaging taken at various time points in a subject’s life, ex-vivo MRI enables us to study how symptoms of disease progression may correlate with changes in brain size. Leveraging sophisticated imaging software, the BRaIN lab also generates 3D image reconstructions of the whole brain from photos taken of individual brain slices at the time of brain removal.
Neurohistology & Neuropathological Diagnostics
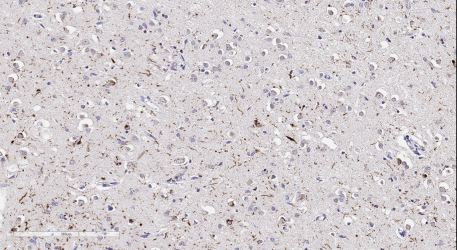
Our fixed research tissue samples are transformed into paraffin-embedded blocks and receive complete neurohistologic work-up, enabling board-certified neuropathologists to provide family members with definitive neuropathological diagnoses and an explanation of symptoms for their loved ones. The BRaIN lab performs immunohistochemical staining for a wide array of markers related to various neurodegenerative processes, aiming to understand the pathologic correlates of brain disease.
Quantitative Neuropathology
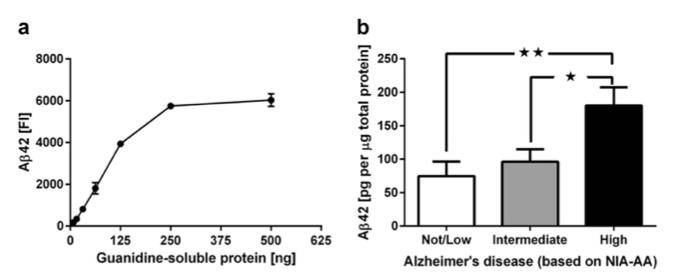
While consensus criteria for the neuropathologic diagnosis of Alzheimer’s disease (AD) rely on ordered rankings of amyloid plaque and neurofibrillary tangle distribution across the brain, these non-quantitative measures have limited utility in correlating pathological features of AD with parameters like age, gender, exposures, genotype, comorbid processes and environmental factors. To provide more robust correlative power, the BRaIN lab performs high-throughput, multiplexed quantitative assays to measure levels of amyloid beta, tau and other toxic proteins across the brain in large patient cohorts. In addition, we leverage AI-driven, high-resolution digital image analysis tools to quantify the distribution and amount of pathology on immunostained slides. These complementary quantitative approaches afford an in-depth view into the molecular drivers of disease.
Citation Policy
Citing REDCap
The UW Medicine Biorepository and Integrated Research (BRaIN) laboratory uses REDCap to store much of the data shared with researchers.
Please use the following boilerplate language in study manuscripts using data provided by the BRaIN Lab.
Study data were collected and managed using REDCap electronic data capture tools hosted at the University of Washington and the Institute of Translational Health Sciences (ITHS)1,2 REDCap (Research Electronic Data Capture) is a secure, web-based software platform designed to support data capture for research studies, providing 1) an intuitive interface for validated data capture; 2) audit trails for tracking data manipulation and export procedures; 3) automated export procedures for seamless data downloads to common statistical packages; and 4) procedures for data integration and interoperability with external sources. The UW and ITHS instance of REDCap is supported by the NCATS/NIH grants UL1 TR002319, KL2 TR002317, and TL1 TR002318..
1 PA Harris, R Taylor, R Thielke, J Payne, N Gonzalez, JG. Conde, Research electronic data capture (REDCap) – A metadata-driven methodology and workflow process for providing translational research informatics support, J Biomed Inform. 2009 Apr;42(2):377-81.
2 PA Harris, R Taylor, BL Minor, V Elliott, M Fernandez, L O’Neal, L McLeod, G Delacqua, F Delacqua, J Kirby, SN Duda, REDCap Consortium, The REDCap consortium: Building an international community of software partners, J Biomed Inform. 2019 May 9 [doi: 10.1016/j.jbi.2019.103208]
